By Santosh Rananaware
Humans are the second most dangerous creatures in the universe. The deadliest creatures in the universe are bacteriophages. Bacteriophages kill more living things everyday than all other organisms combined. Despite their notoriety, however, they do not kill indiscriminately. Bacteriophages are specialized viruses that only kill bacteria.
Bacteria are ubiquitous, unassuming, little microbial organisms that are found everywhere from arctic ice cores to volcanic hot springs. They are also responsible for causing some of the deadliest diseases known to mankind. The Black Plague that ravaged through much of Europe during the 14th century and killed hundreds of millions of people is believed to have been caused by a tiny little bacterium called Yersinia pestis. However, only a very tiny fraction of all the bacteria present in the universe are harmful to any organism at all. In fact, many of them are beneficial to our body and help us in digesting the food we eat.
Fig.1: Enterobacteriophage P4. Illustration Courtesy Ben Darby
Despite their near omnipresence and significant impact on the environment, for much of history humans were oblivious to the existence of bacteria or bacteriophages. This is because of the extremely small size of these creatures. Bacteria are typically only of the size of 1 µm in diameter, which means if you line up a thousand bacteria end-to-end, they would only form a line of about 1 mm long. Bacteriophages are viruses and are a thousand times smaller still. Even today, these minute organisms are visible to us only with the help of extremely strong microscopes.
Despite their infinitesimal size, if you were somehow able to observe these minuscule creatures as they go about their day-to-day activities, you would notice that the bacteria and the phages are in a state of constant war with each other. This war has been going on for centuries. Much of nature’s activities revolve around phages trying to invade bacterial colonies and the bacteria fighting back. Whenever any particular strain of bacteria prosper and threaten to take over a local ecosystem, the phages come like the horsemen of the apocalypse and hunt them down, thereby maintaining balance and diversity.
At this point, it is important to understand that bacteriophages do not ‘hunt’ bacteria in the traditional sense of the word. They do not ‘eat’ their prey like wolves eat deer. Rather, they infect the bacterium with their own genetic material (viral DNA or RNA) and hijack its replication machinery (Fig.2). The bacterium, unaware of the foreign genes being injected in its own genome, replicates them, thereby creating more copies of the virus inside it. After a certain number of viral copies are made and assembled inside the bacterium, it just explodes and releases all those copies into the environment. The released bacteriophages are now free to roam and infect other bacteria.
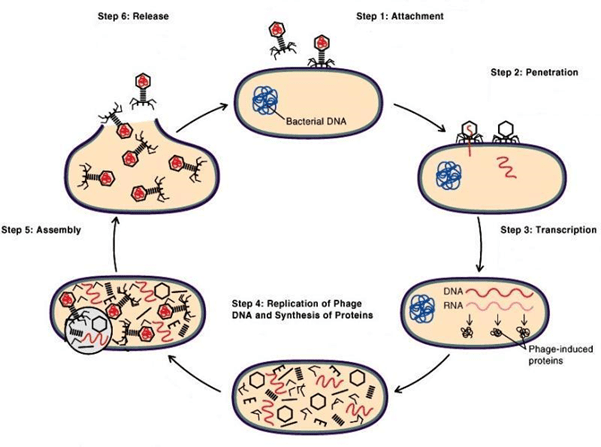
Fig.2: Schematic of bacteriophage replication (Adapted from learn microbiology)
CRISPR/Cas as a bacterial defense mechanism
To counter these phage attacks, the bacteria have evolved to develop many defense mechanisms to protect themselves. For example, one such mechanism adapted by bacteria involves blocking of the phage receptors, which are tiny structures on the surface of bacterial cells that the phages use to attach themselves to the bacteria. By blocking these receptors, the bacteria ensures that the phages are not able to connect to them in the first place and are thereby unable to infect. However, to counter this, the phages have evolved to recognize the blocked receptors and bind to new receptors on the bacterial surface. In this manner, the bacteria and the bacteriophages are in a constant state of evolutionary competition with each other.
Over time the bacteria evolved to develop a sophisticated defense mechanism, known as the CRISPR-Cas system, to protect themselves from the invading phages. The CRISPR-Cas system evolved when bacteria that somehow survived phage attacks started preserving small fragments of the invaders DNA into its own genome at fixed positions in their chromosome called the CRISPR Loci. The CRISPR loci consists of small fragments of DNA from all the different viral invaders that the bacteria was able to survive. These sequence of viral DNA fragments are known as ‘spacer’ sequences and they are sandwiched between ‘repeat’ sequences, which are palindromic sequences of the bacteria’s own DNA that are repeatedly placed between different spacer sequences (Fig.3). The name ‘CRISPR’ is derived from such a peculiar organization of the foreign DNA into the bacterial genome and is the abbreviated form for the term ‘Clustered Regularly Interspaced Palindromic Repeats’.
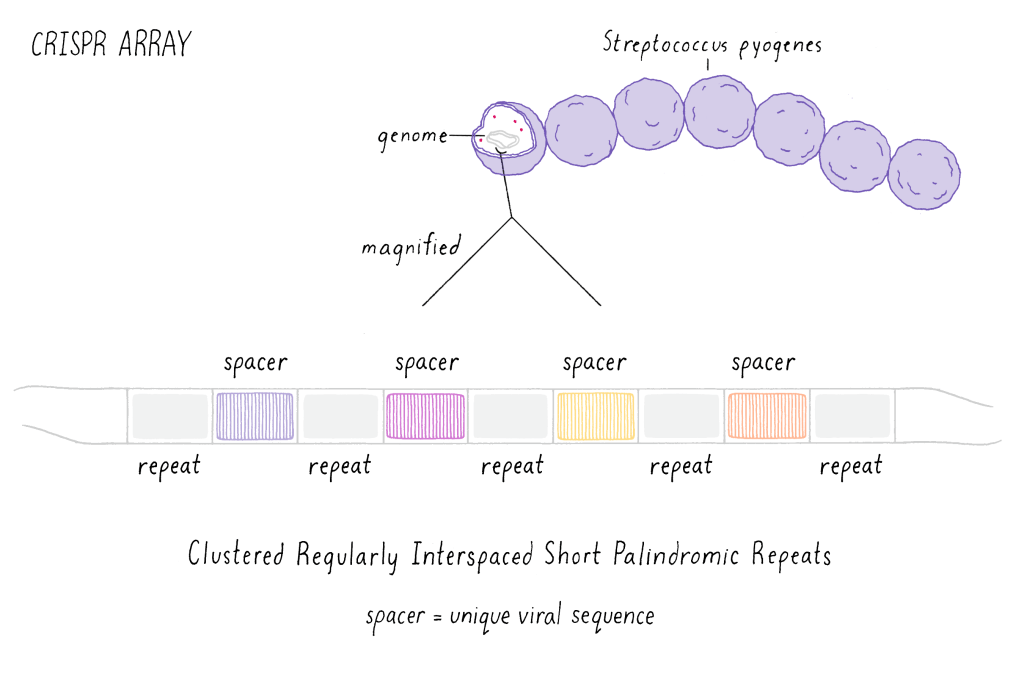
Fig.3: Schematic of spacer and repeat sequences in the CRISPR loci
(Adapted from XBio)
Not much is known about the mechanism by which bacteria acquire and store these spacer sequences. However, Cas1 and Cas2 proteins are believed to play a role in the process. The term ‘Cas’ is the abbreviation for ‘CRISPR associated’ and hence, Cas proteins refer to the family of proteins within the bacteria that are involved in the CRISPR Cas defense mechanism. Every time a new spacer is acquired from a failed bacteriophage attack, it is stored at the leading end of the CRISPR sequence in the bacteria (Fig.4).
The CRISPR loci is eventually transcribed into a short RNA molecule called CRISPR RNA or crRNA. The crRNA then binds with a Cas enzyme which can have one or more nuclease (DNA-cutting) domains that can target and cut any DNA. The crRNA/Cas complex roams around freely inside the bacterial cell. The next time if the same viral bacteriophage invades the bacteria with its genetic material, the crRNA hybridizes with the invading viral DNA through base pairing and guides the Cas enzyme towards it like a homing missile. The Cas enzyme is then able to unwind and cut the foreign DNA, thereby disabling it and protecting the bacteria from its effects.
One might ask that because the spacer sequence stored in the CRISPR array is identical to the foreign DNA injected by the virus, why doesn’t the CRISPR/Cas system accidentally destroy the CRISPR array itself, thinking it to be the invading DNA. The answer to that lies in a short DNA motif that is found only in the invading viral sequence (and not in the spacer stored in the CRISPR array) called the PAM sequence. To prevent potential auto-immunity, a large number of CRISPR/Cas systems in bacteria cleave DNA targets only if they are flanked by short sequences known as the PAM sequences. The spacers present in the CRISPR array lack the PAM sequence and are therefore, immune to CRISPR mediated destruction. The mechanism by which spacers are chosen so that they target only PAM-associated protospacers are still unknown.
The key thing about the CRISPR/Cas defense mechanism, however, is that it is hereditary and hence, subsequent generations of bacteria acquire this phage resistance from their parents without having to have encountered these phages themselves. Around half of all the known bacteria and almost all of archaea are found to have the CRISPR-Cas defense mechanism within them.
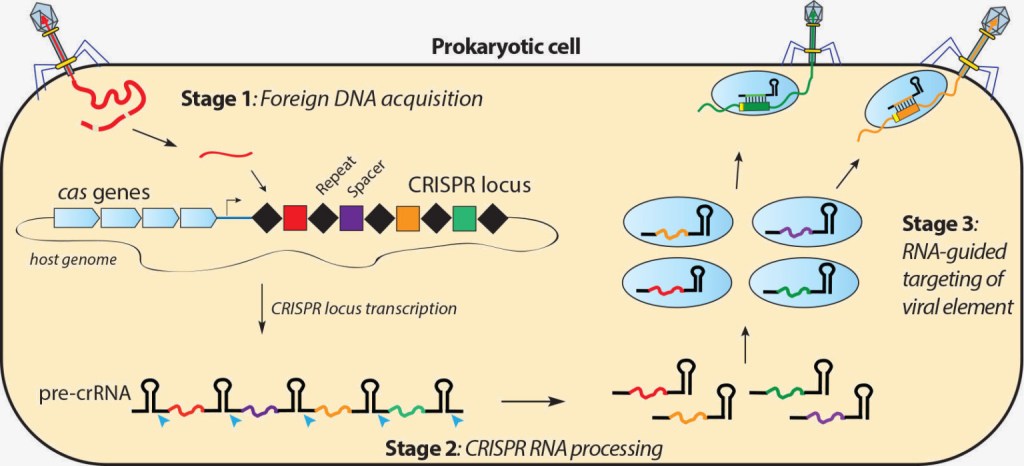
Fig.4: CRISPR/Cas defense mechanism (Reprinted from Doudna Lab)
CRISPR/Cas can be repurposed into a genome editing tool
A key breakthrough in science occurred when scientists realized that the CRISPR/Cas system can be taken out of the bacteria and can be repurposed to easily make precise cuts not just in viral DNA, but in the DNA of any organism! The only components necessary to do so were a guide RNA that can base pair and target the DNA of choice, the presence of a PAM sequence on the target DNA and a Cas enzyme such as Cas9 that can bind and cut the target DNA. Such a system can now be used to target and cut any gene within the genome of any organism. A majority of inheritable diseases, for e.g., sickle cell anemia, occur due to minor genetic defects in the chromosome. The CRISPR/Cas system can now be potentially used to ‘cut’ the defected gene and replace it with a healthy copy of the same gene.
It is amazing to think that a mechanism that evolved in bacteria over thousands of years as a way to protect themselves from their natural enemy, the bacteriophage, could now be harnessed by humans as a tool cut and edit their own DNA.
CRISPR is far from a perfect tool. Scientists have realized that after CRISPR cuts the DNA at the target site, it goes haywire and starts cutting other sites in the DNA thereby leading to unwanted mutations, or off-target effects, in the genome of the target organism. There are also challenges involved delivering the CRISPR/Cas components inside the human cell and increasing its efficiency of on-target editing. A tremendous amount of research is currently ongoing to improve and perfect it.
Meanwhile, however, the bacteriophages (deadliest organisms on earth, if you remember) have evolved to find a way to completely circumnavigate around the CRISPR defense in bacteria. It was recently discovered that the phages produce something that has come to be known as as anti-CRISPR proteins to help them fight CRISPR. Anti-CRISPR proteins are highly diverse but most block CRISPR in one of three ways: (a) Inhibiting DNA binding, (b) Inhibiting crRNA loading, and (c) Inhibiting DNA cleavage. Thus, the arms race continues.